ENA Submission
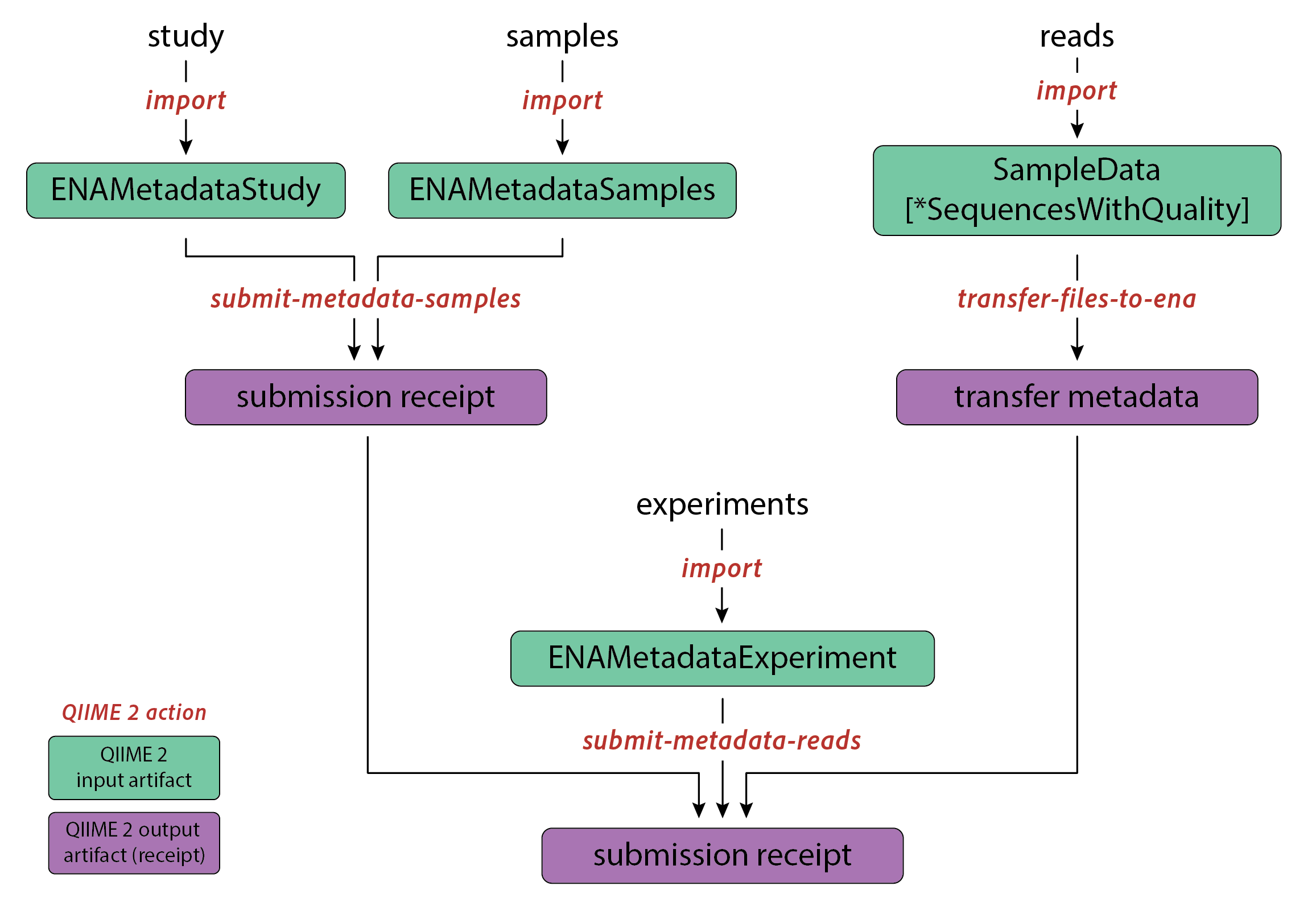
After completing the installation and creating an ENA account, the submission process using the q2-ena-uploader consists of several steps:
It is recommended to create an ENA account and obtain credentials at least 24 hours prior to submission.
Import data into QIIME 2 artifacts.
Upload sample and study metadata to ENA.
Transfer raw reads to the ENA FTP server.
Upload experiment metadata to ENA.
Steps 2-4 should be performed in the specified order. Alternatively, the submit-all action can be used to
submit metadata and raw reads in a single step. To jump to the section on simplified submission, click here.
Step 1: Import data into QIIME 2 artifacts¶
Data imported into QIIME 2 are organized as data artifacts. Each artifact is assigned a Semantic Type, which determines its corresponding Artifact Class within the QIIME 2 framework.
Within our ENA submission workflow:
Raw reads can be imported into QIIME 2 using several different artifact classes, depending on the specific data format. An overview of these supported formats and import choices is available in the official QIIME 2 documentation.
Metadata — including studies, samples, and experiments — are imported as QIIME 2 artifacts with the Semantic Types
ENAMetadataStudy,ENAMetadataSamples, andENAMetadataExperiment, respectively.
These artifacts represent all required inputs for the ENA submission workflow. Once imported, they are stored as .qza files within the QIIME 2 environment. Proceed to the next sections to import all necessary input data into QIIME 2.
Raw reads¶
This is a simple example illustrating the import of FASTQ sequencing data into QIIME 2 artifacts. More import options are described here .
qiime tools import \
--type 'SampleData[PairedEndSequencesWithQuality]' \
--input-path my-sequence-data/ \
--input-format CasavaOneEightSingleLanePerSampleDirFmt \
--output-path demux.qzaThe sections bellow describe all types of metadata imports.
Study¶
To import the metadata of a study into the corresponding QIIME 2 artifacts, run:
qiime tools import \
--type ENAMetadataStudy \
--input-path study_metadata.tsv \
--output-path study_metadata.qzaTo create a valid study TSV file, two mandatory parameters are required: alias and title. All other parameters are optional. When constructing a valid study metadata TSV file, consider consulting one of the provided examples in the Templates section.
Sample¶
To import the sample metadata into the corresponding QIIME 2 artifacts, run:
qiime tools import \
--type ENAMetadataSamples \
--input-path sample_metadata.tsv \
--output-path sample_metadata.qzaChecklists¶
For sample submission, ENA provides metadata checklists detailing the minimal attributes required for different sample types. Please review the full range of ENA sample checklists before submitting your samples.
When using the ENA default sample checklist for all your samples, you do not need to specify the default code ERC000011. However, if you apply multiple checklists for different samples, you must include all relevant ENA sample checklists codes in the metadata column checklist. In this case, create a single metadata TSV file that combines all columns from these checklists and fill in values only where applicable.
Using multiple checklists within the same sample set can be useful when samples originate from different environments, such as gut microbiomes and soil. Conversely, allowing missing values (e.g., for control samples) is appropriate for special cases within a single checklist.
Sample checklist templates are available in this GitHub repository . Each folder contains a sample.tsv file ready for submission. Remember to add an ENA checklist column to indicate which checklist is used. You can also combine columns from multiple checklists in a single submission.
Alternatively, you can download a similar TSV directly from the ENA web portal after logging in and selecting “Register Samples.” Just make sure to add the ENA checklist column and remove the first line of the downloaded file.
Minimal sample structure
Minimal sample structure refers to the ENA default sample checklist data structure. To create a valid TSV file, four mandatory parameters are needed for each sample:
alias(sample ID)taxon_idgeographic location (country and/or sea)collection date
When constructing a valid study metadata TSV file, consider consulting one of the provided examples in the Templates section.
Experiment¶
To import the experiment metadata into the corresponding QIIME 2 artifacts, run:
qiime tools import \
--type ENAMetadataExperiment \
--input-path experiment_metadata.tsv \
--output-path experiment_metadata.qzaFor experiment submission, ENA supports controlled vocabulary in the metadata fields, which can be accessed here.
Minimal experiment structure
The minimal experiment structure requires the following mandatory fields:
titlestudy_ref(study alias or accession number - should correspond to the alias indicated in the study metadata)sample_description(sample ID)platforminstrument_modellibrary_strategylibrary_sourcelibrary_selectionlibrary_layoutlibrary_nominal_length(only for paired reads)library_nominal_sdev(only for paired reads)
The field library_construction_protocol is optional.
When constructing a valid experiment metadata TSV file, consider consulting one of the provided examples, see more in Templates section.
Step 2: Upload sample and study metadata to ENA¶
Set ENA credentials¶
Before uploading to ENA, you need to set two environmental variables containing your ENA credentials:
export ENA_USERNAME=<Webin-XXXXXX>
export ENA_PASSWORD=<password>Please ensure that your credentials were created at least 24 hours before your first submission to the ENA server.
Upload metadata¶
Execute the following QIIME 2 action to submit Study and Sample metadata to perform a test submission to the ENA dev server:
qiime ena-uploader submit-metadata-samples \
--i-study study_metadata.qza \
--i-samples sample_metadata.qza \
--p-action ADD \
--p-dev \
--p-submission-hold-date <hold date> \
--o-submission-receipt samples_receipt.qza--i-study: Artifact containing metadata of the study.--i-samples: Artifact containing metadata of the samples.--p-action-type: 2 action types are supported: ADD (default) and MODIFY.--p-dev: A boolean parameter indicating whether the submission is a test.--p-submission-hold-date: The release date of the study, on which it will become public along with all submitted data. By default, this date is set to two years after the date of submission. Users can specify any date within two years of the current date. Accepted date format isYYYY-MM-DD.--o-submission-receipt: The output artifact containing the assigned ENA accession numbers for the submitted objects.
You can submit a study and sample metadata either separately or together, only one of the corresponding artifacts is required for submission. However, please note that to submit raw reads later, both the study and samples must already exist on the ENA server.
The submission hold date is required for all submissions to the ENA server.
To perform a test submission, set the --p-dev parameter to True (this is also the default). This will submit the data to the ENA dev server
(this data will be removed automatically after 24h). To submit the data to the production server, set the parameter to False or use the --p-no-dev flag.
Step 3: Transfer raw reads to the ENA FTP server¶
Before submitting the Experiment, you must first transfer your files to the ENA FTP server.
Ensure again that your environment variables ENA_USERNAME and ENA_PASSWORD are properly set.
Execute the following QIIME 2 action to transfer the FASTQ files to the ENA FTP server.
qiime ena-uploader transfer-files-to-ena \
--i-demux <your reads artifact> \
--p-action ADD \
--o-metadata transfer_metadata.qza--i-demux: The demultiplexed sequencing data, either single-end or paired-end.--p-action: Specifies the action to take. The default is ADD, but you can use DELETE to remove files from the ENA FTP server.--o-metadata: This is the output artifact containing information about the transfer or deletion status of files on the ENA FTP server.
Step 4: Upload experiment metadata to ENA¶
Make sure that your credentials are configured through the environment variables ENA_USERNAME and ENA_PASSWORD, as described here.
Execute the following QIIME 2 action to submit Experiment/Run metadata to ENA:
qiime ena-uploader submit-metadata-reads \
--i-demux <your reads artifact> \
--i-experiment experiment_metadata.qza \
--i-samples-submission-receipt samples_receipt.qza \
--i-file-transfer-metadata transfer_metadata.qza \
--p-submission-hold-date <hold date> \
--p-action ADD \
--p-dev \
--o-submission-receipt receipt.qza--i-demux: The demultiplexed sequencing data, either single-end or paired-end.--i-experiment: Artifact containing experiments submission parameters.--i-samples-submission-receipt: Artifact containing the submission receipt of the study and samples.--i-file-transfer-metadata: Artifact containing the metadata of the file transfer to the ENA FTP server.--p-action: 2 action types are supported: ADD (default) and MODIFY.--p-dev: A boolean parameter indicating whether the submission is a test.--p-submission-hold-date: The release date for the data submission, determining when it will become public. The accepted date format is YYYY-MM-DD.--submission-receipt: The output artifact containing the assigned ENA accession numbers for the submitted objects.
To perform a test Submission, set the --p-dev parameter to True (this is also the default). This will submit the data to the ENA dev server
(this data will be removed automatically after 24h). To submit the data to the production server, set the parameter to False or use the --p-no-dev flag.